Manoil Lab
MUTANT DISTRIBUTION IS BEING DISCONTINUED. Individual mutant and pooled mutant library distribution will end October 15, 2022 and we will stop accepting new orders September 15, 2022. We will continue to distribute our few remaining complete single mutant arrayed libraries for P. aeruginosa, A. baumannii and K. pneumoniae until December 15, 2022 or until our copies of them are exhausted.
Klebsiella pneumoniae Mutant Library
Individual strains and copies of the Klebsiella pneumoniae Three-Allele Library are available for a charge to the research community through a non-profit cost center at the University of Washington. The library was made in a derivative of the K. pneumoniae strain KPNIH1, an ST258 clinical isolate from a hospital outbreak (http://www.ncbi.nlm.nih.gov/bioproject/73191). The derivative (called MKP103) is deleted of the carbapenemase gene carried on a large conjugal plasmid (pKPQIL) in KPNIH1. The plasmid is otherwise intact. The transposon used for mutagenesis is a derivative of Tn5 carrying a chloramphenicol resistance determinant (transposon T30), and mutants were generated using transposon-transposase complex (“transposome”) mutagenesis.
Please be aware that the library parent strain, MKP103, is itself chloramphenicol resistant. In our hands, the MIC of MKP103 on chloramphenicol is roughly 50mg/mL. Mutants were selected on a much higher concentration of chloramphenicol, 175mg/mL. For more information, please see Ramage, et al. 2017. J. Bacteriol. 199:e00352
The library has 2-3 separate insertions per gene, and strains have been colony purified and re-sequenced. There are approximately 12,000 mutants in the library, corresponding to ~85% of the predicted KPNIH1 genes. Most of the unrepresented genes presumably correspond to those essential for growth on the medium used to generate the mutants (LB containing 175mg/mL chloramphenicol). The strains have been single colony-purified, and the insertion locations for most of them have been confirmed by re-sequencing. Individual mutants are available, as is the library parent strain (MKP103) and the wild-type (KPNIH1).
Requesting individual mutants
The library Excel file (which can be downloaded from the link below) details information about each mutant in the three-allele library. Most of the column headings are self-explanatory. There is also a description of column headings on the second sheet of the Excel file (“Legend”).
To request individual mutants, fill out the order form that corresponds to your type of institution (academic or nonacademic; links below) and send the completed Excel order form to kpmutant [ at ] uw.edu. There is an instruction sheet in the order form file, but please let us know if you have any questions about the form itself, the mutants, or the ordering process.
For requests from countries requiring import or other permits, the requestor must obtain the necessary permits and email copies to kpmutant [ a t ] uw.edu. All charges resulting from failure to provide the required permits will be paid by the requestor, including the cost of return shipment following customs rejection. While email is preferred, if it is necessary to fax copies of permits, they may be faxed to Manoil lab at +1-206-685-7301.
We are able to accept payment by purchase order number, check, bank transfer, or credit card for smaller orders. Credit card payments may be made via a secure website; the link to the website will be provided with the invoice. We require a purchase order to be emailed with the order form for orders exceeding $1,000.
Choice of Strains
In creating a large arrayed mutant library like this one, it is inevitable that some assignments will fail to check out. We have done our best to minimize cross-contamination and insertion mis-assignment by colony purification and two rounds of sequencing. In addition, high throughput growth and distribution may lead to some mixed cultures.
We have included multiple independent insertion mutants for most genes, and suggest that multiple mutants corresponding to genes of interest be requested to help provide coverage in case individual mutants cannot be confirmed. In quality control tests of the three-allele library, a small percentage of mutant insertion sites did not match the original assignments. Unfortunately, we are unable to provide replacements for mutants that cannot be confirmed. We also recommend using the parent to this library (MKP103) for comparisons.
Receipt and maintenance of strains
Individual strains are sent as stab cultures in semisolid agar. Samples of the strains received should be maintained as frozen stocks (–80°C) in the recipient laboratory. We recommend that the researcher streak from the stab onto a nutrient medium such as LB agar (without antibiotic) immediately after receipt. After overnight growth scoop up a generous sample from the dense part of the streak for the frozen stock (in LB containing 5% DMSO).
Every strain is viable at the time of shipping. It is possible that a strain you requested is viable for only a short period of time due to the mutation it harbors, and would not be recoverable after shipping. Once a strain has been shipped, there will be no refunds or reshipments without additional orders.
Confirmation of Strains
We urge investigators to check the identities of mutants by PCR or sequencing prior to use, and to share this information with us for incorporation into the strain database. We recommend you test 10 individual colonies by PCR both with flanking primers and with a transposon-specific primer paired with a flanking primer to confirm your mutants.
For each gene of interest, the researcher should design appropriate flanking primers. These primers should be initially tested using the wild type parent strain. To show that the intact gene is absent in the insertion strain, PCR with the same primers should yield either no band or a band corresponding to a much larger product.
To show the presence of the transposon insertion, use a transposon-specific primer with one of the flanking primers. For transposon T30, use transposon-specific primer Pcm-140 (5’-CTGCGAAGTGATCTTCCGTCAC-3'). The flanking primer should be chosen according to the orientation of the transposon relative to the insertion site. For ‘F’ (forward) insertions, the transposon-specific primer will point back toward lower genome positions.
When you have either positively or negatively confirmed a mutant strain, please send an email reporting your result to kpmutant [at] uw.edu.
Ordering Library Copies
In making replicates of the entire library for distribution, several steps of quality control are performed. Strains are assessed for their growth by visual assessment of turbidity and strains that did not grow well are grown up individually and included in supplemental plates.
Library orders may be initiated either by contacting us at kpmutant [at] uw.edu or by filling out the Excel order form (instructions are on the first sheet) and emailing the form to kpmutant [at] uw.edu.
Please contact us with any questions about these products and the ordering process.
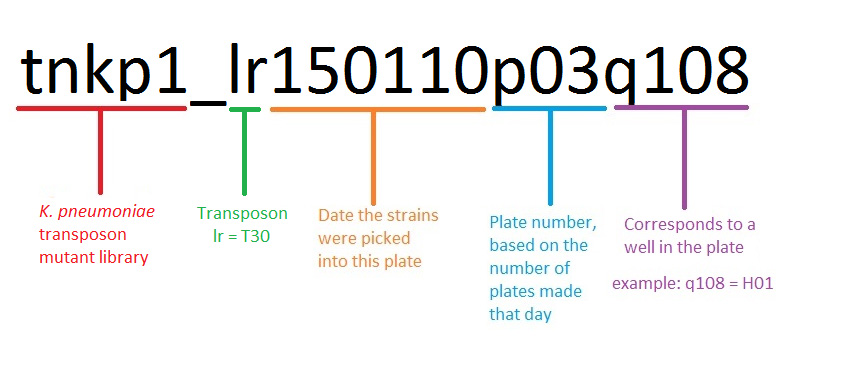
Mutant Naming
In publications, please reference strains from the three-allele Library by Strain Name/ Location (unique identifier) and refer to the genotype in the following way: gene name (or locus if there is no gene name)-well name (final three digits of the strain name) as the allele number::Transposon name (T30). For example, for strain tnkp1_lr150110p03q105, the genotype is hisD105::T30, and for straintnkp1_lr150110p03q101, the genotype is KPNIH1_00420-101::T30.
Please follow these links for order forms and additional information
Academic Institution order form
Non-Academic Institution order form
library